Updated 30 September 2025 (c) 2025
Introduction
In the past few decades, modern science has uncovered a universe that is far vaster and more awe-inspiring than ever imagined before, together with a set of elegant natural laws that deeply resonate with the idea of a cosmic lawgiver. Along this line, 46% of Americans, including, interestingly enough, 54% of atheists, 55% of agnostics and 43% of nones, say that they experience a “deep sense of wonder about the universe” on at least a weekly basis [Masci2016].
In spite of these exhilarating developments, some writers, principally of the creationist and intelligent design schools, prefer instead a highly combative approach to science, particularly to traditional topics such as geology and evolution. One widely used line by such writers is that certain features of biology are so unlikely, according to simple back-of-the-envelope probability calculations, that they could never have been produced by a purely natural, “random” evolutionary process, even assuming millions of years of geologic history. Thus the entirety of evolutionary theory is proved mathematically false. Why don’t scientists see the light?
For example, some writers equate the theory of evolution to the absurd suggestion that monkeys randomly typing at a typewriter could compose a selection from the works of Shakespeare, or that an explosion in an aerospace equipment yard could produce a working airliner [Dembski1998; Foster1991; Hoyle1981; Lennox2009]. More recent studies of this genre argue that functional biology operates on an exceedingly small subset of the space of all possible DNA sequences, and that any changes to the “computer program” of biology are, like changes to human computer programs, almost certain to make the organism non-functional [Axe2017; Marks2017].
One typical creationist-intelligent design argument addresses the human alpha-globin molecule, a component of hemoglobin that performs a key oxygen transfer function in blood. These writers argue that since alpha-globin is a protein chain based on a sequence of 141 amino acids, and since there are 20 different amino acids common in living systems, the “probability” of selecting human alpha-globin “at random” is one in 20141, which is roughly one in 10183 (i.e., a one followed by 183 zeroes). This probability is so tiny, so they argue, that even after millions of years of random molecular trials, no human alpha-globin protein molecule would ever appear, thus refuting the hypothesis of human evolution [Foster1991, pg. 79-83; Hoyle1981, pg. 1-20; Lennox2009, pg. 163-173].
The treacherous world of probability and statistics
While not generally appreciated by the public at large, it is a well-known fact in the world of scientific research that arguments based on probability and statistics are fraught with numerous potential fallacies and errors, so that a very strict methodology is required: What exactly is the phenomenon being modeled? What exactly is the probability space (the set of all possible outcomes)? How exactly is probability to be measured? Is each possible event presumed to have the same probability? If so, why can this be assumed? Are certain events assumed to be independent? Is so, why can this be assumed? If these and other questions are not carefully addressed, then it matters not in the slightest how sophisticated the mathematical calculations are — the chain of inference is broken, and any conclusion is almost certainly invalid.
One overriding lesson of probability and statistics, when rigorously applied, is that seemingly improbable “coincidences” can and do happen. For instance, a common classroom exercise is to inquire how likely it is, in a class say of 30 students, that two or more of the students have the same birthday. Most people presume this is very unlikely, but the correct probability is 70.6%. For additional details and examples, see [Hand2014; Mlodinow2009; Pinker2021].
Fundamental fallacies in creationist-intelligent design probability arguments
So how do the creationist-intelligent design probability arguments rate by these standards? To begin with, these writers seldom if ever explicitly define the underlying probability space, model or measure, so they are in serious trouble right from the start. For example, when they say “the probability of selecting human alpha-globin at random,” they appear to be asking for the probability of selecting human alpha globin based on an all-at-once random assemblage of atoms. But this all-at-once construction is most assuredly not the scientific hypothesis of how they formed. Instead, available evidence from many published studies indicates that human alpha-globin arose as the end product of a long sequence of intermediate steps, each of which was biologically useful in an earlier context. See, for example, the article [Hardison2012], which cites 144 papers on the topic of hemoglobin evolution.
More importantly, when these writers reckon a probability as merely the reciprocal of the total number of theoretical possibilities, they implicitly presume that every instance, say of the theoretical space of 141-long amino acid sequences, is equally likely to actually arise in a biological species such as Homo sapiens. But no justification is given for this sweeping equiprobable assumption, and it is certainly false — some amino acid sequences are relatively likely to emerge, while vast numbers of other sequences are not biologically possible at all. No empirically defensible statistics are known for such phenomena (we have only one example of human evolution), so to assume that all such events are equiprobable is invalid and misleading.
Furthermore, most anti-evolution probability arguments (certainly including the alpha-globin example) fail to recognize that the process of natural biological evolution is not really a “random” process. Evolution certainly has some random aspects, notably mutations and genetic events during reproduction. But the process of natural selection, acting in a competitive landscape and with numerous complicated environmental pressures, is anything but random. This strongly directional nature of natural selection, which is the essence of evolution, by itself invalidates many anti-evolution probability arguments.
In short, most creationist-intelligent design probability arguments presume an utterly unrealistic probability model, and this presumption completely invalidates their conclusions. For additional details, see Jason Rosenhouse’s comments on the “argument from improbability” in his newly published book on the topic [Rosenhouse2022].
Extraterrestrial aliens made this rock
To illustrate some of these difficulties, consider this example:
While out hiking, I found this rock. The following table gives measurements made on the rock. The first two rows give the overall length and width of the rock. Each of the next six rows, after the first two, gives thickness measurements, made on a 3cm x 6cm grid of points from the top surface. All measurements are in millimeters:
| Measurement or row | Column 1 | Column 2 | Column 3 |
| Length | 105.0 | ||
| Width | 48.21 | ||
| Row 1 | 35.44 | 35.38 | 36.54 |
| Row 2 | 38.06 | 38.27 | 38.55 |
| Row 3 | 38.02 | 39.53 | 39.29 |
| Row 4 | 38.66 | 40.50 | 41.96 |
| Row 5 | 39.40 | 43.48 | 43.31 |
| Row 6 | 39.58 | 41.83 | 43.07 |
This is a set of 20 measurements, each given to four significant figures, for a total of 80 digits. Among all rocks of roughly this size, the “probability” of a rock appearing with this particular set of measurements is thus one in 1080. Note that this probability is so remote that even if the surfaces of each of the ten planets estimated to orbit each of 100 billion stars in the Milky Way were examined in detail, and this were repeated for each of the estimated 100 billion galaxies in the visible universe, it is still exceedingly unlikely that a rock with this exact set of measurements would ever be found. Thus this rock could not have appeared naturally, and must have been created by space aliens…
Wait a minute! It is just an ordinary rock!
What is the fallacy in the above argument? First of all, modeling this set of measurements as a random variable of 80 digits, and assuming all measurements are equiprobable, is an erroneous reckoning. In real rocks, the measurement at one point is constrained by physics and geology to be reasonably close to that of nearby points. Presuming that every instance in the space of 1080 theoretical digit strings is equally probable as a set of rock measurements is unjustified and invalid, as we have noted above. Thus the above reckoning must be rejected on this basis alone.
The post-hoc probability fallacy
More importantly, any reckoning of this type falls prey to the post-hoc probability fallacy, which is a form of confirmation bias reasoning — specifying a probability test after the fact, based on specific measurements that have already been performed, and then claiming a remarkable result. This is equivalent to dealing a nondescript ordered hand of cards, such as the one to the right, calculating its probability after the fact (it is approximately one in 4 x 1021), and then claiming that it could not have occurred naturally — the dealer must have cheated? In reality, of course, there is nothing particularly remarkable about this hand of cards at all. The post-hoc probability calculation is completely misleading.Key point: It is important to note that the post-hoc fallacy does not just weaken a probability or statistical argument. In most cases, the post-hoc fallacy completely nullifies the argument. The correct reckoning is “What is the probability of X occurring, given that X has been observed to occur,” which of course is unity. In other words, the laws of probability, when correctly applied to a post-hoc phenomenon, can say nothing one way or the other about the likelihood of the event.
In general, probability reckonings based largely on enumerations of theoretical possibilities have no credibility when applied to real-world problems. And reckonings where a statistical test or probability calculation is specified after the fact are completely invalid and must be immediately rejected — some outcome had to occur, and the laws of probability by themselves cannot help to determine its likelihood.
The post-hoc fallacy in creationist-intelligent design writings
As mentioned above, one typical creationist-intelligent argument is that since human alpha-globin is a protein chain based on a sequence of 141 amino acids, and since there are 20 different amino acids common in living systems, the “probability” of selecting human alpha-globin at random is one in 20141, or one in approximately 10183. This probability is so tiny, so these authors argue, that even after millions of years of random molecular trials, no human alpha-globin protein molecule would ever appear.
But this line of reasoning is a dead-ringer for the post-hoc probability fallacy. Note that this probability reckoning was performed after the fact, not on any new data but on a single very limited dataset (the human alpha-globin sequence) that has been known in the biology literature for decades. Some sequence had to occur, and the fact that the particular sequence found today in humans was the end result of the course of evolution provides no guidance, by itself, as to the probability of its occurrence.
Further, just like the rock measurements above, the enumeration of theoretical possibilities in this calculation has no credibility. Some alpha-globin sequences are likely to be realized while others are biologically impossible. It may well be that there is nothing particularly special about the human alpha-globin sequence, as evidenced, for example, by the great variety in alpha-globin molecules seen across the biological kingdom, all of which perform a similar oxygen transfer function. But there is no way to know for sure, since we have only one example of human evolution. Thus the reckoning employed here, namely enumerating theoretical possibilities, assuming all possibilities are equiprobable, and taking the reciprocal to obtain a probability figure, is invalid and misleading.
How scientific research fields deal with the post-hoc fallacy
It is worth noting here that many other fields of scientific research have recognized the great dangers of the post-hoc fallacy and confirmation bias reckoning, and have instituted strict methodologies to avoid them. Some examples include:
-
Preregistration: Prior to starting a study, researchers publicly announce their plans, including all data analysis and statistical testing methods. This procedure prevents after-the-fact tinkering, or selectively reporting certain analyses, or changing the statistical methods until they obtain the finding they hypothesized. This has now been adopted in numerous fields, ranging from pharmaceutical trials to the social sciences [Nesi2024].
-
Data blinding: As data is collected, a “blind” (a pseudorandomly generated value) is added to each data point. Then only after the researchers have gathered all their data, finished their statistical analysis and even written the first draft of their report, the computer subtracts the blinds to reveal the true results. This has now been adopted in numerous large-scale particle physics, astronomy and cosmology research projects [Caldwell2024; Savitsky2025].
-
Data holdout: In mathematical finance and some related fields, the post-hoc fallacy is known as “backtest overfitting.” Thus it is now common practice to use only part of a historical dataset when developing a model, preserving the other part to finally evaluate the model. However, it is very important not to repeat this process, since repeated usage of the holdout data is itself subject to the post-hoc fallacy [Bailey2021].
Needless to say, creationist and intelligent design writers do not adopt highly rigorous procedures of this sort to avoid the post-hoc fallacy. Indeed, most do not even seem to be aware of the potential problems here.
Hemoglobin and chlorophyll
Also with regards to alpha-globin, it is worth noting that heme, the key oxygen-carrying component of hemoglobin, is remarkably similar to chlorophyll, the molecule behind photosynthesis. The principal difference is that heme has a central iron atom, whereas chlorophyll has a central magnesium atom; otherwise they are virtually identical. This similarity can hardly be a coincidence, and in fact researchers concluded since at least 1980, based on both functional and biochemical evidence, that these two biomolecules “have arisen in the course of evolution from a common origin” [Hendry1980; Hardison2012]. Here is a diagram of the two molecules [from MasterOrganicChemistry.com]:
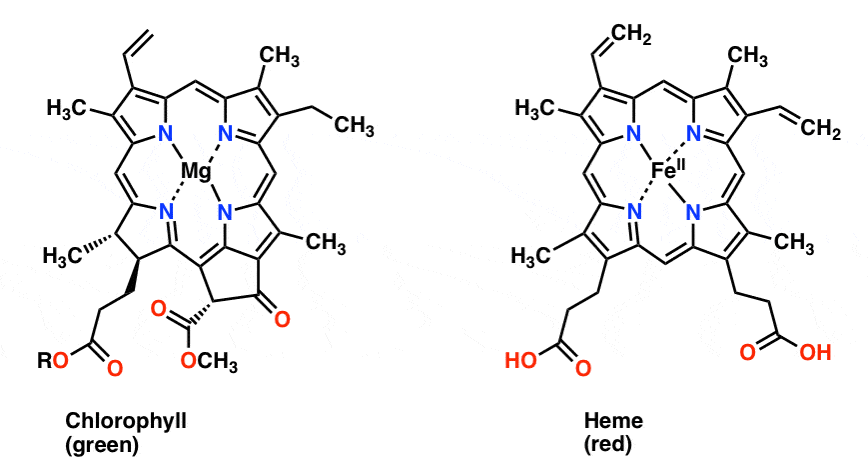
Summary of probability fallacies
In summary, the probability arguments that have been promoted in the creationist and intelligent design literature are riddled by severe errors, most notably that they are based on utterly unrealistic probability models and are clear instances of the post-hoc probability fallacy. Far from being decisive arguments against evolution, instead they are empty arithmetic exercises with no relevance to real empirical biology. Such arguments would never be accepted in a rigorously peer-reviewed journal in evolutionary biology or applied probability, not because of their implication for evolution (pro or con), but because this type of reasoning is well-known to be invalid in numerous other contexts. For additional discussion, see [Musgrave1998; Rosenhouse2016; Rosenhouse2018]. A newly published book by Jason Rosenhouse, which discusses these issues in significantly greater detail, is [Rosenhouse2022].
Computer programs emulating biological evolution
If there were some underlying mathematical or biological principle that forbids biological structures from evolving novel features, surely this principle should be seen in computer simulations of evolution. But quite the opposite is true — such studies abundantly confirm that evolutionary processes really do work, not just in biology but in other arenas as well.
For example, as mentioned above, some critics have equated natural biological evolution to the absurd suggestion that some monkeys typing randomly at a keyboard could generate a passage of Shakespeare. Others have argued that any changes to a “computer program” would surely render the program unusable. But these too are fallacious arguments, since they ignore the all-important process of natural selection. As a single example, a 2009 study by the present author exhibited results of a computer program simulating natural evolution, which “evolved” segments of English text very much akin to actual passages from Charles Dickens. In many instances, a class of college students were unable to distinguish the computer-generated text segments from real text segments taken from Dickens’ Great Expectations. See English-text or [Bailey2009] for details.
Another example is the recent rise of “genetic algorithms” and “evolutionary computing,” namely computer programs that mimic the process of biological evolution to produce novel solutions to scientific and engineering problems. As a single example, in 2017 Google researchers generated 1000 image recognition algorithms, each of which were trained using state-of-the-art deep neural networks to recognize a selected set of images. They then used an array of 250 computers, each running two algorithms, to identify an image. Only the algorithm that scored higher proceeded to the next iteration, where it was changed somewhat, mimicking mutations in biological evolution. The researchers found that their scheme could achieve accuracies as high as 94.6%, better than human efforts [Gershgorn2017]. In another Google-funded research project, a computer was programmed with only the rules of Go, together with an evolution-style “deep learning” algorithm, and then had the program play games against itself. Within a few days it had advanced to the point that it defeated an earlier Google program 100 games to zero. This earlier program, in turn, had previously defeated the world’s champion human Go player [Greenmeier2017]. As a third example, researchers in Spain have developed software that employs an evolution-like strategy to directly infer scientific results from raw data, in some cases more successfully than the best human efforts [Wood2022].
Some creationist writers have attempted to dismiss examples of genetic algorithms and evolutionary computing, claiming that such programs always include a target that nullifies their result. But this is not true, certainly not for the above-mentioned examples [Thomas2006; Thomas2010].
Novel features in nature
It is worth emphasizing here that nature can and often does produce remarkable structures and features, by the well-understood evolutionary processes of mutation, shuffling of genes and natural selection. Here are just a few of many examples that could be mentioned:
-
Lenski’s 2012 E. coli experiment: In January 2012, a research team led by Richard Lenski at Michigan State University demonstrated that colonies of viruses can evolve a new trait in as little as 15 days. The researchers studied a virus, known as “lambda,” which infects only the bacterium E. coli. They engineered a strain of E. coli that had almost none of the molecules that this virus normally attaches to, then released them into the virus colony. In 24 of 96 separate experimental lines, the viruses evolved a strain that enabled them to attach to E. coli, using a new molecule that they had never before been observed to utilize. All of the successful runs utilized essentially the same set of four distinct mutations [Zimmer2012; Lenski2022].
-
Synthesis of RNA nucleotides and other biomolecules: Many scientists hypothesize that RNA (a molecule similar to DNA, with four “bases” or nucleotides) was involved in the origin of life (see Origin). As recently as 1999, the appearance of RNA nucleotides on the primitive Earth was widely thought to be a “near miracle” by researchers in the field [Joyce1999]. Nonetheless, in May 2009 a team led by John Sutherland of the University of Manchester discovered a particular combination of chemicals, very likely to have been plentiful on the early Earth, that synthesized the RNA nucleotides cytosine and uracil, which are known as the pyrimidines [Wade2009]. More recently, in 2018, Carell’s team demonstrated a single process that created all four nucleotides [Service2018]. In short, the natural production of the four RNA nucleotides, once thought to be “impossible,” is now fairly well understood.
-
Hawaiian crickets. In the 1800s, a species of chirping crickets was introduced to the Hawaiian Islands, where they became quite common. However, the Hawaiian crickets have a fearsome predator — dive-bombing flies that target chirping crickets, then implant their larvae in them. In the 1990s, researchers noted that a field in Kauai that previously was the home to many crickets now seemed silent. However, a nighttime search found that in fact there were lots of crickets there, but very few of the males now chirped — in just five years, or roughly 20 generations, a mutation had arisen that inhibited many of the males from chirping, thus protecting the population [Zuk2013, pg. 81-82].
-
Recent human evolution. A 2010 study found a total of 30 recently evolved genetic variants that are distinct among natives of Tibetan highlands, permitting them to live well at very high altitudes (more efficient metabolism, avoid overproducing red blood cells in thin air, etc.). They concluded that these changes constitute the fastest documented case of human evolution [Wade2010b]. Some other instances of recent human evolution include: (a) a genetic adaptation has arisen among Southern Chinese that makes them more resistant to alcohol, at the cost of turning red in the face; (b) gene changes in Eskimo populations that help them better cope with bitter cold; (c) genetic changes among certain primitive farming people that enhance vitamin B9 production, which is absent in their tuber diets; (d) two genetic mechanisms for lighter skin color, which among higher-latitude people promotes vitamin D production — Europeans have one, while East Asians have another; and (e) a gene that promotes hair with thicker shafts among East Asians, possibly as added protection against cold [Wade2010c].
-
Evolution of the coronavirus. The Covid-19 pandemic of 2020-2024, which as of September 2024 has claimed the lives of over 18,000,000 worldwide, including over 1,000,000 Americans, has underscored the reality of evolutionary novelty in a way that affects every person on the planet. The rise of the delta and omicron variants of the virus (the latest is “KP.3”), which are significantly more transmissible than earlier variants, is a particularly striking case of “evolution before our eyes.” No one “designed” these deadly virus variants [Achenbach2021; Adam2022; Schumaker2021; Zimmer2023].
Is independent creation a credible alternative?
Does the hypothesis of independent creation of individual species (i.e., with no common biological ancestry), as advocated by many creationist and intelligent design writers, provide a reasonable alternative in terms of probability?
Here it is instructive to consider transposons or “jumping genes,” namely sections of DNA that have been “copied” from one part of an organism’s genome and “pasted” seemingly at random in other locations. The human genome, for example, has over four million individual transposons in over 800 families [Mills2007]. In most cases transposons do no harm, because they “land” in an unused section of DNA; and some transposons have subsequently adopted biological functionality (although for the purposes of this discussion it does not matter in the least whether or not they have biological functionality). But because they are distinctive and inherited, they serve as excellent markers for genetic studies. Indeed, transposons have been used to classify a large number of vertebrate species into a family tree, with a result that is virtually identical to what biologists had earlier reckoned based only physical features and biological functions [Rogers2011, pg. 25-31, 86-92]. As just one example, consider the following table, where columns labeled ABCDE denote five blocks of transposons, and x and o denote that the block is present or absent in the genome [Rogers2011, pg. 89].
Transposon blocks
Species A B C D E
/-------- Human o x x x x
/--------- Bonobo x x x x x
/ \-------- Chimp x x x x x
/----------- Gorilla o o x x x
-----|----------- Orangutan o o o x x
\----------- Gibbon o o o o o
It is clear from these data that our closest primate relatives are chimpanzees and bonobos. As another example, here is a classification of four cetaceans (ocean mammals) based on transposon data [Rogers2011, pg. 27]:
Transposon blocks
Species A B C D E F G H I J K L M N O P
/------ Bottlenose dolphin x x x x x x x x x x x x x x x x
/\------ Narwhal whale x x x x x x x x x x x x x x x x
---|------- Sperm whale x x x x x o o o o o o o o o o o
\------- Humpback whale x x o o o o o o o o o o o o o o
Other examples could be listed, encompassing an even broader range of species [Rogers2011, pg. 25-31, 86-92].
Needless to say, these data, which all but scream “descent from common ancestors,” are highly problematic for creationists and others who hold that the individual species were separately created without common biological ancestry. Transposons typically are several thousand DNA base pair letters long, but, since there are often some disagreements from species to species, let us be very conservative and say only 1000 base pair letters long. Then for two species to share even one transposon starting at the same spot, presumably only due to random single-letter mutations since creation, the probability (according to the creationist hypothesis, and assuming equiprobable mutations) is one in 41000 or roughly one in 10600. For 16 such common transposons, the chances are one in 416000 or roughly one in 109600 (and individual species typically have many thousands of transposons).
But this is not all, because we have not yet considered the fact that in each diagram above, or in other tables of real biological transposon data, there is a clear hierarchical relationship. This is by no means assured, and in fact is quite improbable — for almost all tables of “random” data, there is no hierarchical pattern, and no way to the rearrange the rows to be in a hierarchical pattern. For example, in a computer run programmed by the present author, each column of the above cetacean table was pseudorandomly shuffled (thus maintaining the same number of x and o in each column), and the program checked whether the rows of the resulting table could be rearranged to be in a hierarchical order. There were no successes in 10,000,000 trials. As a second experiment, a 4 x 16 table of pseudorandom data (with a 50-50 chance of x or o) was generated, and then the program attempted to rearrange the rows to be in a hierarchical pattern as before. There were only three successes in 10,000,000 trials.
These calculations are simplified and informal; more careful reckonings can be done, and one can vary the underlying assumptions. But one way or the other, it is clear that the hypothesis of separate creation of individual species does not resolve any probability paradoxes; instead it enormously magnifies them.
Some creationists propose that a Supreme Being separately created individual species with hundreds of thousands of transposons already in place, essentially just as we see them today. But since this hypothesis can be crafted to match any set of DNA evidence, it fails to be falsifiable and thus fails to be truly scientific. More importantly, this presumption merely replaces a scientific failure (the inability of the independent creation model to explain, using natural scientific laws, the vast phylogenetic patterns in transposon data) with a theological disaster (why did a truth-loving Supreme Being fill the genomes of the biological kingdom with vast amounts of misleading DNA evidence, if “descent from common ancestors” is not the conclusion we are to draw?) [Rogers2011, pg. 89]. Indeed, with regards to the discomfort some have about evolution, the creationist-intelligent design alternative of separate creation is arguably much worse, both scientifically and theologically.
Probability, statistics and evolution done right
It should be added here that researchers in evolutionary biology, phylogenetics, applied mathematics and computer science have for many years applied rigorous advanced probability and statistical techniques, typically based on DNA sequence data, to infer the evolutionary history of various elements of the biological kingdom. This is a rapidly growing field with many researchers and numerous remarkable results. Some good references for those who wish to learn more about these methods include: [Bromham2016; Warnow2017; Wiley2011].
Conclusions
It is certainly true that there are numerous complex structures in biology; alpha-globin is but one of many thousands that could be listed. How did these structures first arise, and how did they evolve to the forms we see today? For that matter, was the origin of life on the early Earth an inevitable development, or was it an improbable event unlikely to have been repeated anywhere in Milky Way galaxy? Researchers in these fields would be the first to acknowledge that there is still much that is not yet fully understood. For example, leading researchers often disagree as to whether the origin of the key biochemical components of life was inevitable or unlikely. This is a large, dynamic field with many researchers worldwide, spanning disciplines from organic chemistry and DNA to astronomy and astrobiology, and with thousands of peer-reviewed research papers published each year.
However, as we have shown above, the simple back-of-the-envelope probability arguments that have appeared in the creationist-intelligent design literature do not help unravel these profound questions, because these arguments are riddled with severe errors that invalidate their conclusions and would disqualify them from rigorously peer-reviewed journals in evolutionary biology or applied probability, not because of their implication for evolution (pro or con), but instead because such reasoning is well-known to be invalid in numerous other contexts. These difficulties include:
-
Presuming an utterly unrealistic probability model, such as presuming that all instances of a 141-long amino acid sequence are equally likely to occur in human alpha-globin, so that the “probability” of any particular instance is merely the reciprocal of the total number of theoretical possibilities. Such reckonings, based on enumerating theoretical possibilities rather than real empirical data, have no credibility.
-
Calculating the probability of an event, such as the appearance of the human alpha-globin sequence, after the fact, and then claiming it could not happen naturally. This is a clear instance of the post-hoc probability fallacy, analogous to dealing a nondescript hand of cards from a well-shuffled deck, calculating its probability after the fact, and then claiming that this event could not have happened naturally. The post-hoc fallacy (a form of confirmation bias reasoning) by itself nullifies many creationist-intelligent design probability arguments.
-
Implicitly presuming that a biomolecular structure came into existence “at random” via a chance assemblage of atoms. To the contrary, abundant evidence shows that such structures are the result of a long series of intermediate steps over the eons, each useful in an earlier context.
-
Relying on sophisticated mathematical analyses and calculations, but ignoring the fact that since the underlying probability model is an invalid description of the phenomenon in question, it does not matter in the slightest how sophisticated these mathematical calculations are.
-
Ignoring the fact that a very wide range of biomolecules could perform the essential functions of the given biomolecule, so that the odds given against the formation of the given biomolecule are highly exaggerated.
-
Ignoring the fact that biological evolution is fundamentally not a “random” process — mutations may be random, but natural selection, the essence of evolution, is far from random.
-
Ignoring a large body of published studies showing that evolution can and often does produce remarkable structures and features.
Perhaps at some time in the distant future, a super-powerful computer will be able simulate with convincing fidelity the multi-billion-year biological history of the Earth, in the same way that scientists today attempt to simulate, in a much more modest scope, the Earth’s weather and climate. Then, after thousands of such simulations have been performed, researchers might obtain some meaningful statistics on the chances involved in the origin of life on Earth, or in the formation of some class of biological structures such as alpha-globin. Perhaps also researchers will eventually reconstruct, in the laboratory, additional key biomolecular steps involved in the origin and early development of life. And perhaps one day researchers will even discover forms of life on other planets, and eventually, after thousands of such life forms have been catalogued, they may be able to empirically assess the probability of the origin of life or of specific biomolecules. Until that time, the probability calculations that appear in creationist-intelligent design literature should be viewed with great skepticism, to say the least.
After all, does anyone really believe that the entire edifice of modern evolutionary biology — hundreds of thousands of peer-reviewed published papers — can be felled by a few deeply fallacious back-of-the-envelope probability calculations that do not involve any new empirical data? Common sense says otherwise. As mathematician Jason Rosenhouse writes [Rosenhouse2018],
When biologists ascribe to evolution the ability to craft information-rich genomes, they are neither speculating nor guessing. The basic components of evolutionary theory are empirical facts. Genes really do mutate, sometimes leading to new functionalities. The process of gene duplication with subsequent divergence leads to the creation of information by any reasonable definition of the terms. Selection can string small variations together into directional change. On a small scale, this has all been observed. And if small increases in information are an empirical reality on human timescales, then what abstract principle of mathematics is going to rule out much larger increases on geological scales?
Then here come the ID [intelligent design] folks, full of swagger and bravado. They say the accumulated empirical evidence must yield before their back-of-the-envelope probability calculations and abstract mathematical modeling. Evolution should be abandoned in favor of the new theory of intelligent design. This theory states, in its entirety, that an intelligent agent of unspecified motives and abilities did something at some point in natural history. Not very useful.
In a larger context, it is not clear that highly technical issues such as biomolecular structures or probability calculations have any place in a discussion of modern philosophy or theology. Further, such intellectual acrobatics are not even necessary: As noted above, the natural universe, as revealed by the latest state-of-the-art scientific research, is far larger, far more exotic, far more astounding in complexity and beauty, and, and the same time, far more bound by elegant laws, than any previous generation could ever have imagined. And the fact that we humans, working together across races, economic classes and national boundaries, can uncover these laws, via diligent application of the scientific method, is a tribute to the highest potential of the human spirit.

